Outcome from NMFk analysis. (A) The Silhouette-Reconstruction
Download scientific diagram | Outcome from NMFk analysis. (A) The Silhouette-Reconstruction criterium (see the Methods Section). On the x-axis is denoted the number of basis lipid configurations and on the y-axis-the average Silhouettes (right y-axis, red marking) as from publication: Unsupervised Machine Learning for Analysis of Coexisting Lipid Phases and Domain Growth in Biological Membranes | Phase separation in mixed lipid systems has been extensively studied both experimentally and theoretically because of its biological importance. A detailed description of such complex systems undoubtedly requires novel mathematical frameworks that are capable to decompose and | Unsupervised Machine Learning, Lipids and Biological Membranes | ResearchGate, the professional network for scientists.

Average reconstruction error (red) and the average Silhouette width

Unsupervised Machine Learning for Analysis of Coexisting Lipid Phases and Domain Growth in Biological Membranes

Outcome from NMFk analysis. (A) The Silhouette-Reconstruction criterium

GitHub - SmartTensors/NMFk.jl: Nonnegative Matrix Factorization + k-means clustering and physics constraints for Unsupervised and Physics-Informed Machine Learning
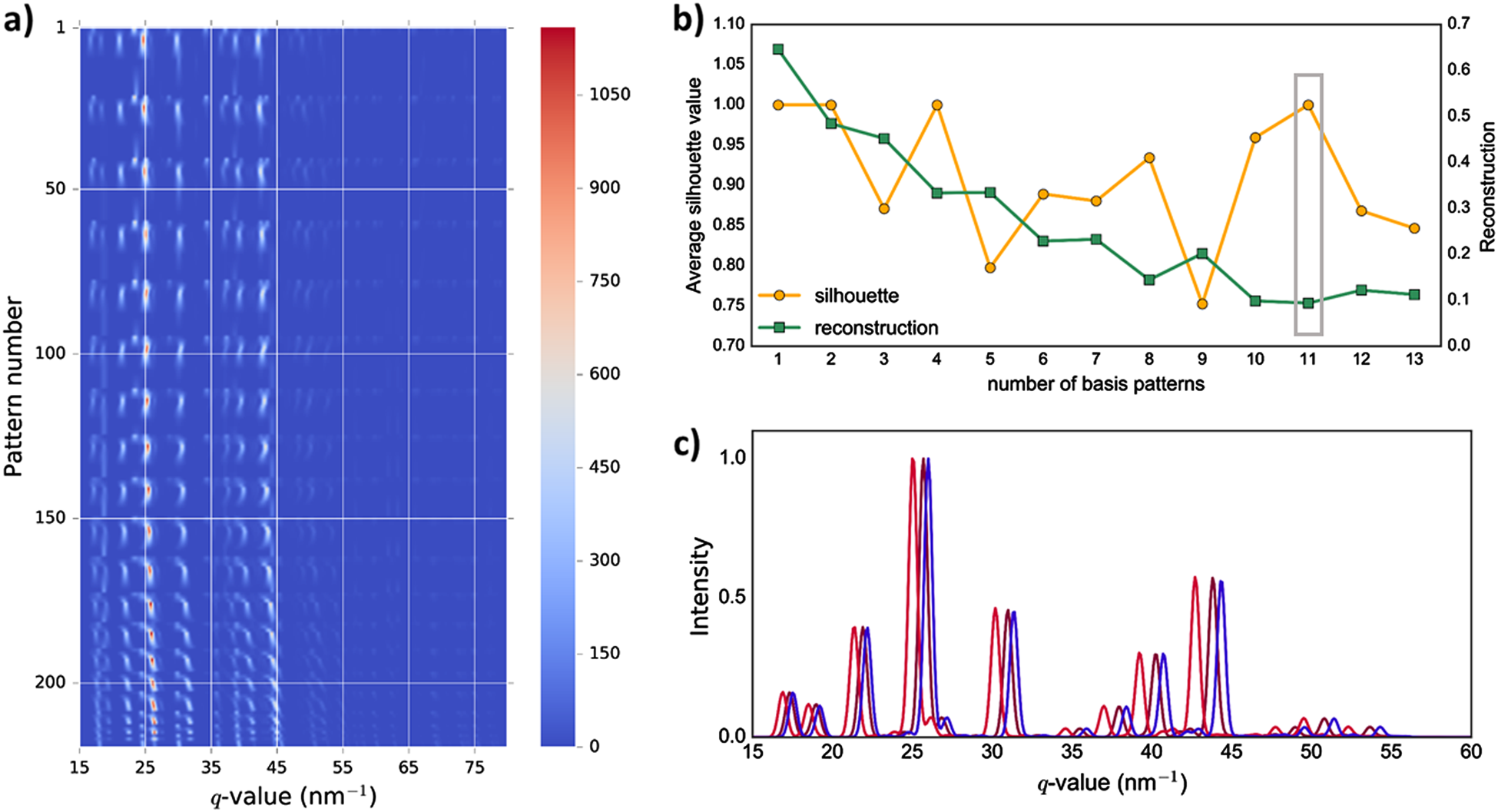
Unsupervised phase mapping of X-ray diffraction data by nonnegative matrix factorization integrated with custom clustering

Unsupervised Machine Learning for Analysis of Coexisting Lipid Phases and Domain Growth in Biological Membranes

AIC and Silhouette values computed for an example X matrix using NMFk

k Scaling for clustering and Silhouette—The runtimes of clustering and

PDF] Towards Accurate 3D Human Body Reconstruction from Silhouettes

a) Silhouette values from clustering (♢) and NMF reconstruction error

Average reconstruction error (black) and the average Silhouette width









