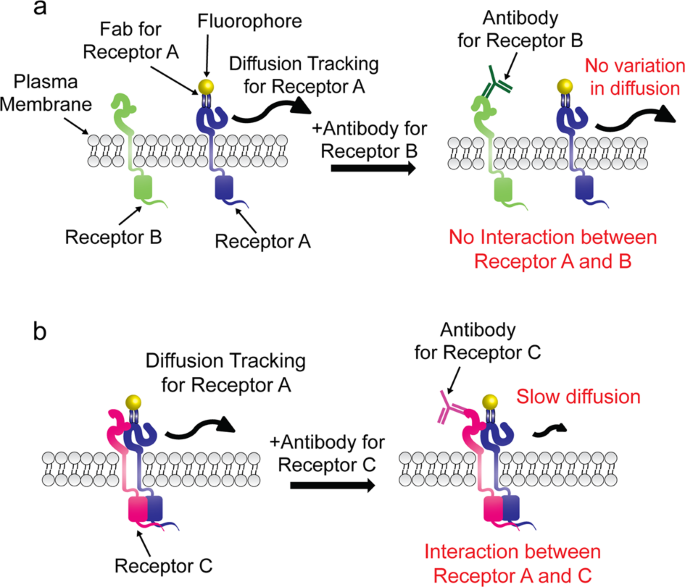
Analysis of transient membrane protein interactions by single-molecule diffusional mobility shift assay | Experimental & Molecular Medicine

Frontiers | A Gene-Based Positive Selection Detection Approach to Identify Vaccine Candidates Using Toxoplasma gondii as a Test Case Protozoan Pathogen
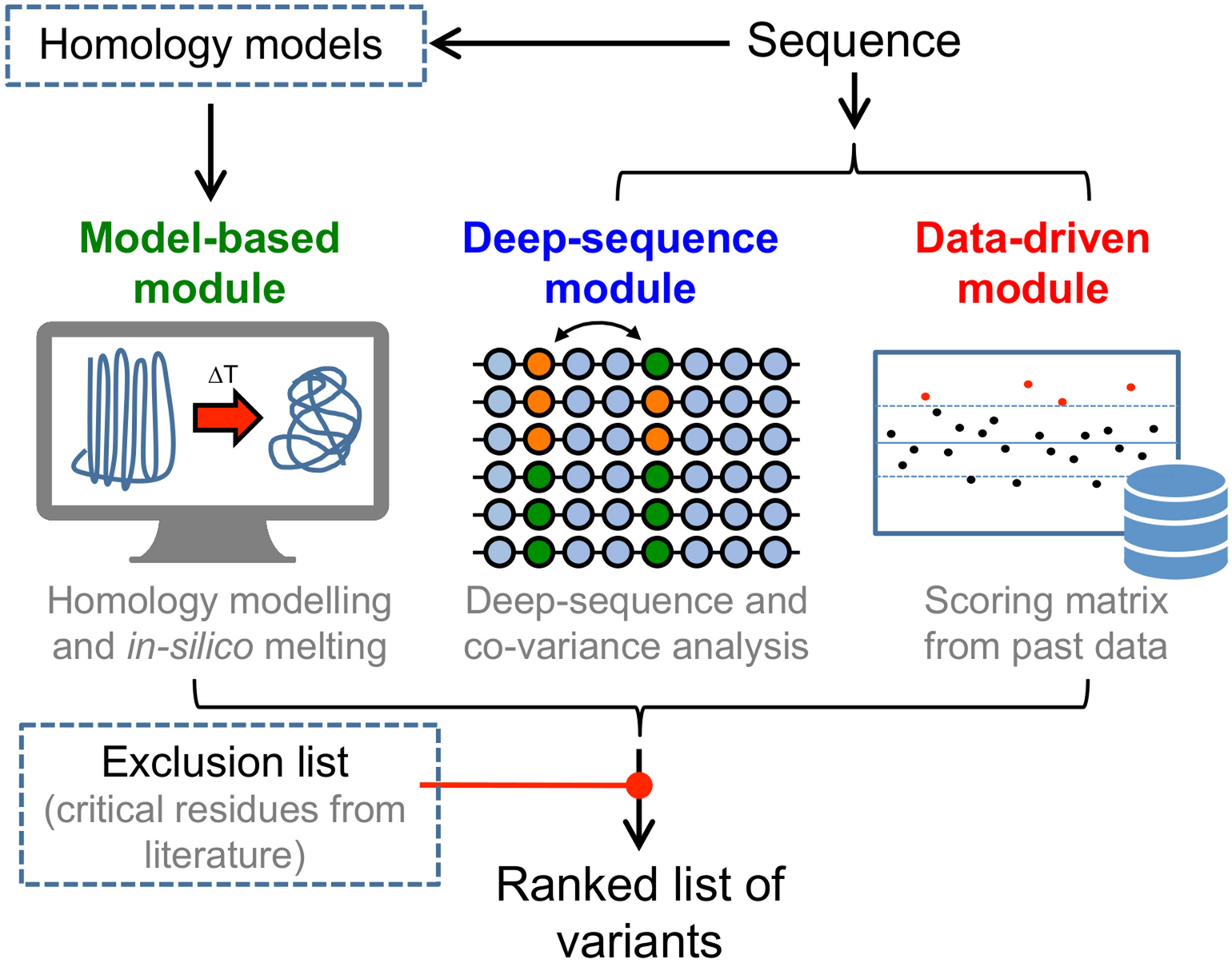
Membrane protein contact and structure prediction using co-evolution in conjunction with machine learning | PLOS ONE

Interplay between hydrophobicity and the positive-inside rule in determining membrane-protein topology | PNAS
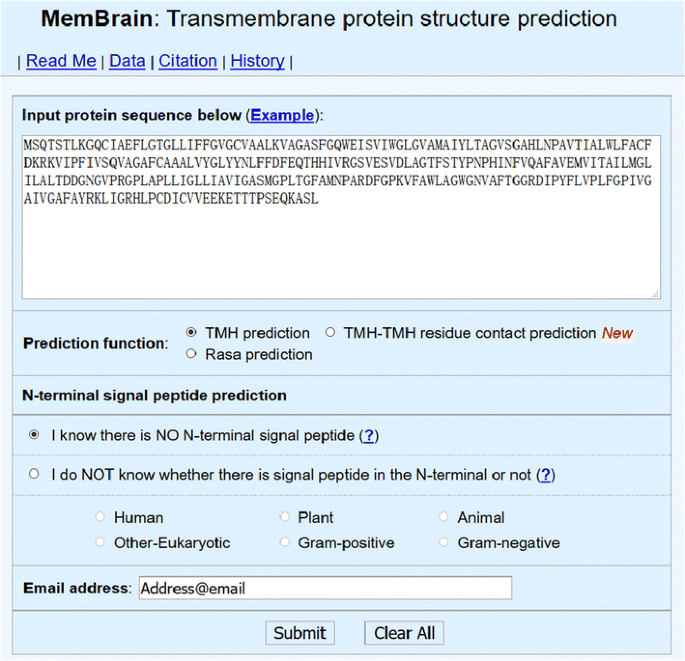
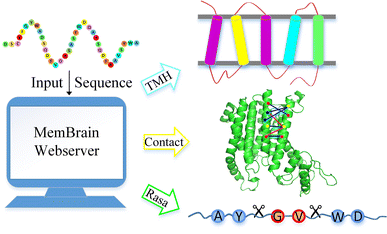
MemBrain: An Easy-to-Use Online Webserver for Transmembrane Protein Structure Prediction | SpringerLink

Multiple C2 domains and transmembrane region proteins (MCTPs) tether membranes at plasmodesmata | EMBO reports
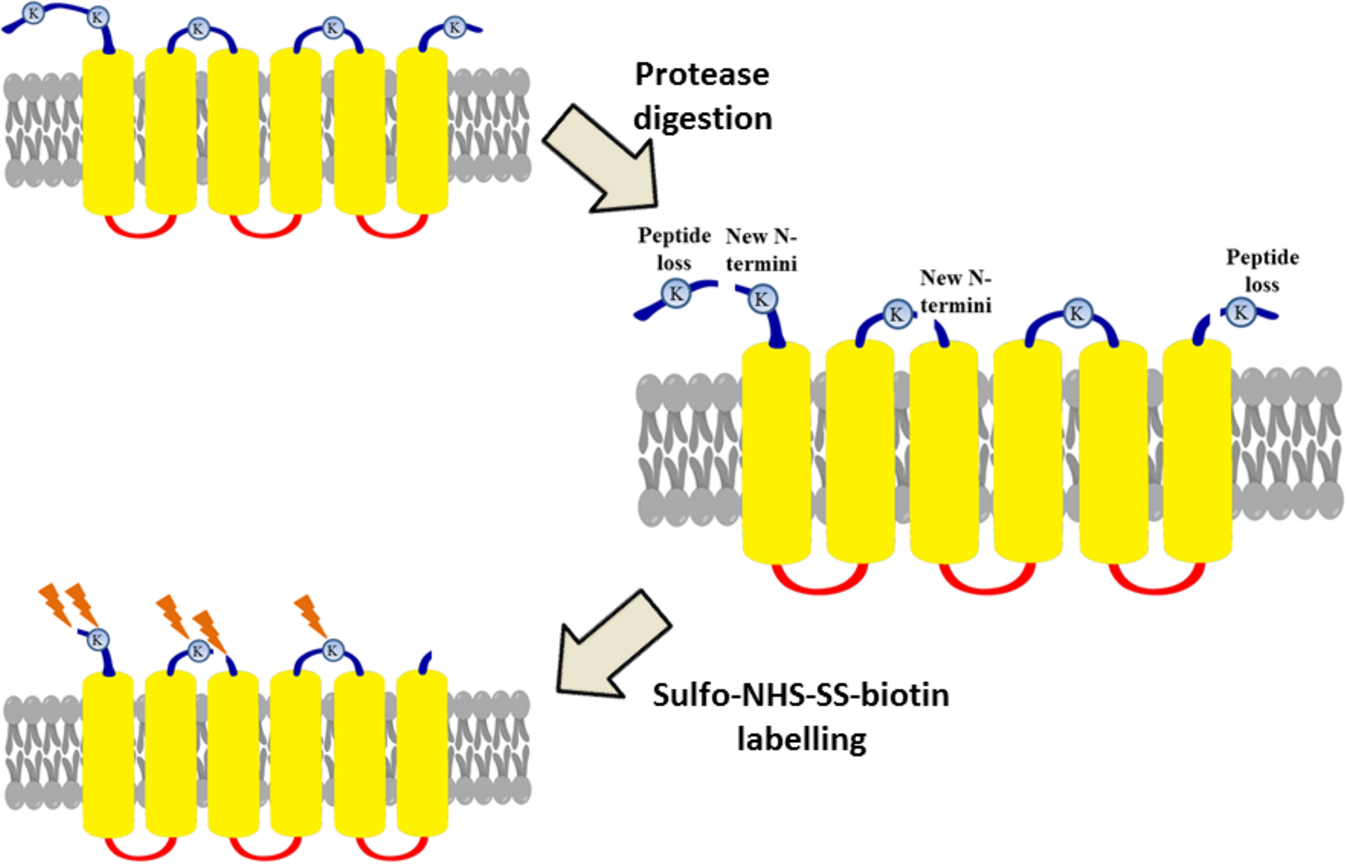
Partial proteolysis improves the identification of the extracellular segments of transmembrane proteins by surface biotinylation | Scientific Reports

Interplay between hydrophobicity and the positive-inside rule in determining membrane-protein topology | PNAS

Methods for Systematic Identification of Membrane Proteins for Specific Capture of Cancer-Derived Extracellular Vesicles - ScienceDirect
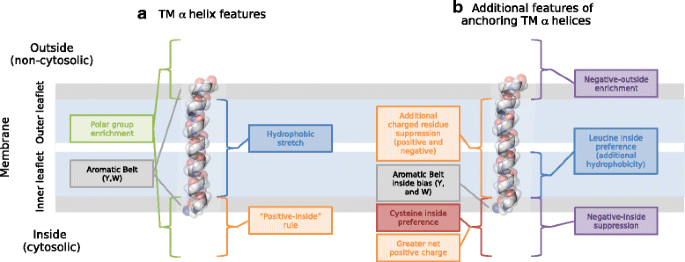
Charged residues next to transmembrane regions revisited: “Positive-inside rule” is complemented by the “negative inside depletion/outside enrichment rule” | BMC Biology | Full Text

MemBrain: An Easy-to-Use Online Webserver for Transmembrane Protein Structure Prediction | SpringerLink
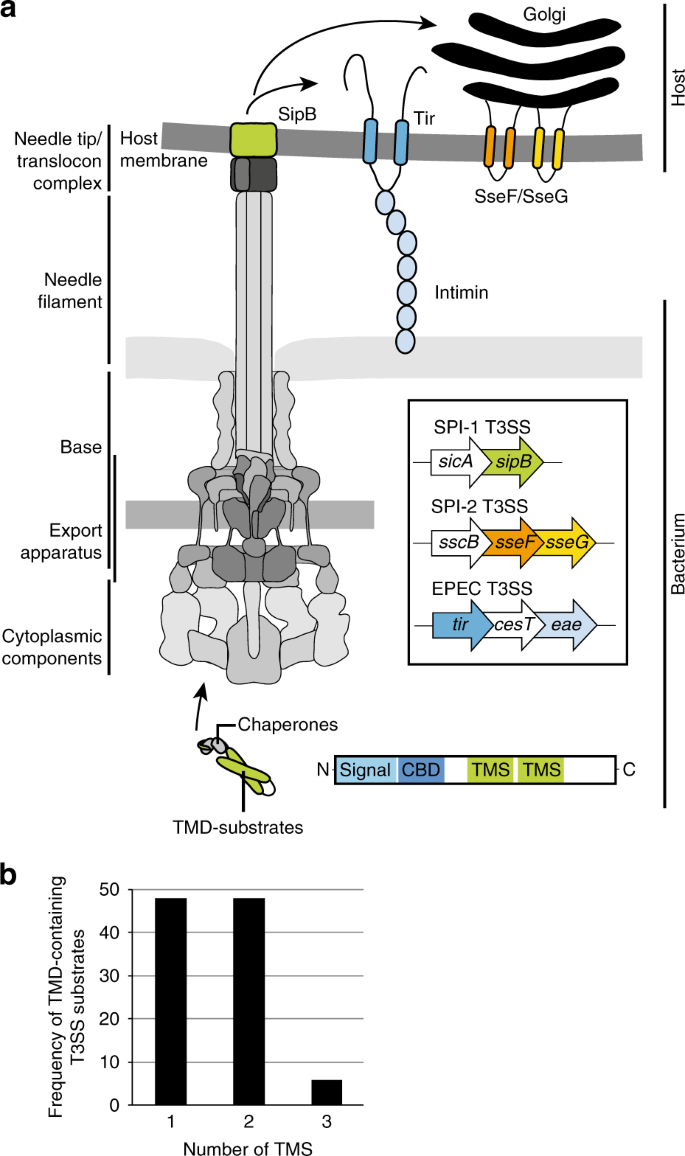
Revealing the mechanisms of membrane protein export by virulence-associated bacterial secretion systems | Nature Communications

Interplay between hydrophobicity and the positive-inside rule in determining membrane-protein topology | PNAS
Membrane protein contact and structure prediction using co-evolution in conjunction with machine learning | PLOS ONE

Interplay between hydrophobicity and the positive-inside rule in determining membrane-protein topology | PNAS
Membrane protein contact and structure prediction using co-evolution in conjunction with machine learning | PLOS ONE